Research
Mitochondrial protein biogenesis
Mitochondria are essential organelles in eukaryotes and best known
as the powerhouse of our cells: they convert the energy from our food into the cellular energy currency ATP. Moreover, mitochondria constitute a central platform in cellular metabolism controlling important biochemical pathways of e.g. amino acids or lipids. They synthesize important cofactors like heme or iron-sulfur clusters and are checkpoints for the cellular suicide program, called apoptosis.
Due to this central role in cell life and death, already slightest impairments of mitochondrial functions can lead to severe human diseases. Therefore, it has to be ensured that all proteins, which mediate these functions are located at the correct position within the organelle.
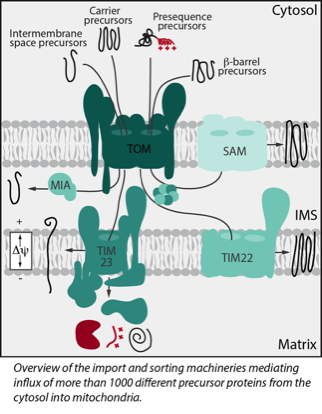
Mitochondria are equipped with more than 1000 different proteins. Most of these proteins are encoded in the nucleus and imported after their synthesis as precursors in the cytosol. Import, sorting into the respective mitochondrial subcompartment and processing of these precursor proteins is mediated by sophisticated protein machineries (see cartoon).
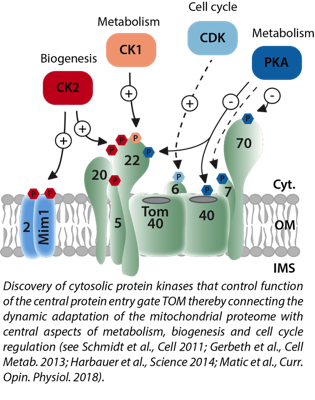
Protein quality control and mitochondrial stress - Signalling at the TOM and MIM protein import machineries
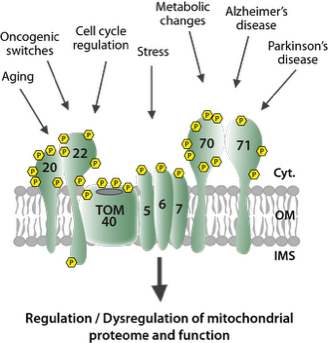
Mitochondria are equipped with more than 1000 different proteins. Most of these proteins are encoded in the nucleus and imported after their synthesis as precursors in the cytosol. Import, sorting into the respective mitochondrial subcompartment and processing of these precursor proteins is mediated by sophisticated protein machineries (see cartoon) Our group studies how the mitochondrial proteome is built, maintained and dynamically adapted upon changing cellular demands. A major focus is on the discovery of cellular signalling mechanisms which dynamically modulate function of the protein import machineries, thereby adapting the mitochondrial proteome to e.g. metabolic switches, stress conditions or disease related changes like oncogenic switches of amyloid-beta toxicity. We also investigate changes in cytosolic signalling networks triggered upon defects in protein import that result in mitochondrial stress.
The mitochondrial protein entry gate TOM as a central hub in controlling metabolic fitness. We aim to discover novel principles and regulatory mechanisms that control metabolic fitness under physiological and pathophysiological conditions.
Chris Meisinger
Institute for Biochemistry and Molecular Biology
Stefan-Meier-Straße 17
D-79104 Freiburg
0049 (0)761 2035287
chris.meisinger(at)biochemie.uni-freiburg.de